Evaluating coverage bias in next-generation sequencing of
By A Mystery Man Writer
Whole-genome sequencing is essential to many facets of infectious disease research. However, technical limitations such as bias in coverage and tagmentation, and difficulties characterising genomic regions with extreme GC content have created significant obstacles in its use. Illumina has claimed that the recently released DNA Prep library preparation kit, formerly known as Nextera Flex, overcomes some of these limitations. This study aimed to assess bias in coverage, tagmentation, GC content, average fragment size distribution, and de novo assembly quality using both the Nextera XT and DNA Prep kits from Illumina. When performing whole-genome sequencing on Escherichia coli and where coverage bias is the main concern, the DNA Prep kit may provide higher quality results; though de novo assembly quality, tagmentation bias and GC content related bias are unlikely to improve. Based on these results, laboratories with existing workflows based on Nextera XT would see minor benefits in transitioning to the DNA Prep kit if they were primarily studying organisms with neutral GC content.

Phables: from fragmented assemblies to high-quality bacteriophage
Genome assembly contig count versus total length of assembly. Each

PDF] Comparison of the sequencing bias of currently available

PDF] Summarizing and correcting the GC content bias in high

PDF] Comparison of the sequencing bias of currently available

Lack of genomic coverage in dfrA-thyA genes reveals deletions in

Depth of Coverage, Non-metric Mulitdimensional Scaling plot. Plot

of serotype predictions that lack an O antigen call by the

The dependence of coverage on GC content. The coverage across

Effect of blood sample preservation agent on DNA yield.

PDF] Summarizing and correcting the GC content bias in high
- TURN YOUR FOUNDATION FROM LOW TO HIGH COVERAGE

- Women Lace Briefs,Triangle Panties,Breathable Thin Briefs,Low Waist Thongs,Half Coverage Panties,Seamless Briefs,Women Night Underwear,5-Pack

- Experts report record low Great Lakes ice coverage

- Why Do I Have Low Signal Strength On My Phone?

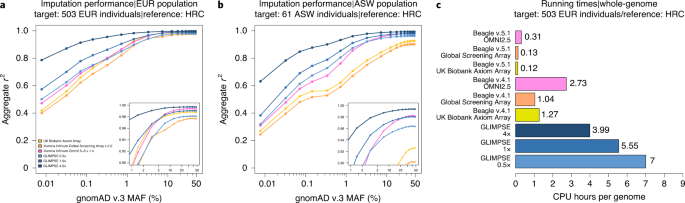
- Efficient phasing and imputation of low-coverage sequencing data
- Side Pockets Blue Springs
- Women's Glamorise 1246 Elegance Front Close T-Back Wonderwire Bra (White 36F)

- Leopard Print Push Thin Bra Set Push High Elastic Intimates - Temu

- Pirate the Maldivian tiger shark. Yoga Leggings are a benefit for Na – OneOceanDesigns

- NIT-Mulheres Glitter Mini Saia De Cintura Alta Discoteca De Lantejoulas Roupas De Desempenho De Palco

