How do I increase the minimum dot size in Seurat's DotPlot function? · Issue #2297 · satijalab/seurat · GitHub

By A Mystery Man Writer
Thank you very much for your hard work in developing the very effective and user friendly package Seurat. I want to use the DotPlot function to visualise the expression of some genes across clusters. However when the expression of a gene
FeatureScatter normalized values below 0 · Issue #2525 · satijalab/seurat · GitHub
DotPlot: cluster order and subsets · Issue #711 · satijalab/seurat · GitHub

How do I increase the minimum dot size in Seurat's DotPlot function? · Issue #2297 · satijalab/seurat · GitHub
How is average expression done in DotPlot? · Issue #825 · satijalab/seurat · GitHub
DotPlot not showing average expression with only one Identity factor · Issue #2583 · satijalab/seurat · GitHub

DotPlot split.by Average Expression in Legend? · Issue #1541 · satijalab/ seurat · GitHub
Expression Values in DotPlot Function in Seurat · Issue #783 · satijalab/ seurat · GitHub

How to add average expression scale to dotplot of merged gene list (plotted onto single dot plot) · Issue #4544 · satijalab/seurat · GitHub

Different percentage of detected cells between DotPlot and FindMarkers function · Issue #5579 · satijalab/seurat · GitHub

Dot plot scaling issue/request · Issue #396 · scverse/scanpy · GitHub
Importance of 'scale' in DotPlot · Issue #5742 · satijalab/seurat · GitHub

DimPlot() changes point shape for more than 50 000 points. · Issue #3897 · satijalab/seurat · GitHub

Average expression of gene on dotplot · Issue #2649 · satijalab/seurat · GitHub

Changing titles of feature gene lists on DotPlots · Issue #5265 · satijalab/ seurat · GitHub
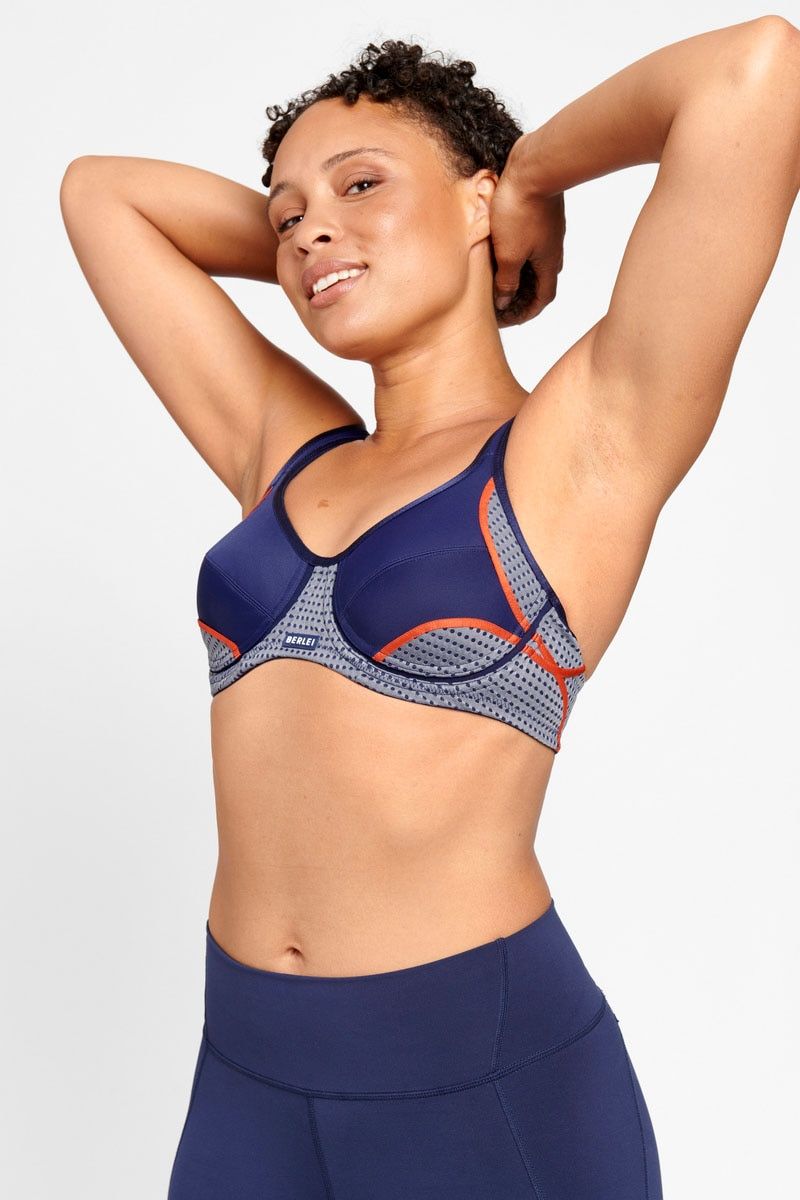
- BERLEI Electrify Underwire Bra

- La Número 1 - Liquidación de invierno❄☔ Casaca para mujer Ahora S/ 65.00🌟 antes S/ 219.00 Hasta 70% de DSCTO en todas las prendas de temporada🔥 Adquiérelo en: Somos La Número
- BKLD 2024 Fashion Velvet Pants Women Autumn Winter Flare Pants High Waist Trousers Street Style Pants Bottoms Women Flare Pants

- 2.Kohl Kohl Jar and Stick Period: New Kingdom Dynasty: Dynasty 18, early Reign: reign of Thutmose I…

- SWAG Boxer Briefs Mens Underwear DORITOS MEDIUM 32 - 34 BLUE COOL RANCH NWT